
conv2
Perform two-dimensional convolution
Syntax
C = conv2(A,B)
C = conv2(hcol,hrow,A)
C = conv2(...,shape)
Description
C = conv2(A,B) performs the two-dimensional convolution of matrices A and B, returning the result in the output matrix C. The size in each dimension of C is equal to the sum of the corresponding dimensions of the input matrices minus one. That is, if the size of A is [ma,mb] and the size of B is [mb,nb], then the size of C is [ma+mb-1,na+nb-1].
C = conv2(hcol,hrow,A) convolves A separably with hcol in the column
direction and hrow in the row direction. hcol and hrow are both vectors.
C = conv2(...,shape) returns a subsection of the two-dimensional convolution, as specified by the shape parameter. shape is a string with one of these values:
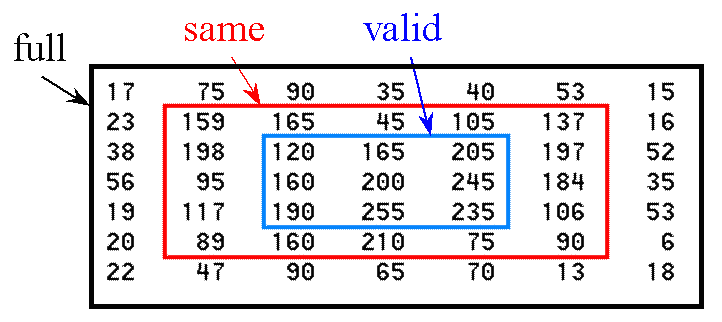
'full' (the default) returns the full two-dimensional convolution.'same' returns the central part of the convolution of the same size as A. 'valid' returns only those parts of the convolution that are computed without the zero-padded edges. Using this option, size(C) = [ma-mb+1,
na-nb+1] when size(A) > size(B).For image filtering, A should be the image matrix and B should be the filter (convolution kernel) if the shape parameter is 'same' or 'valid'. If the shape parameter is 'full', the order does not matter, because full convolution is commutative.
Class Support
All vector and matrix inputs to conv2 can be of class double or of any integer class. The output matrix C is of class double.
Remarks
conv2 is a function in MATLAB.
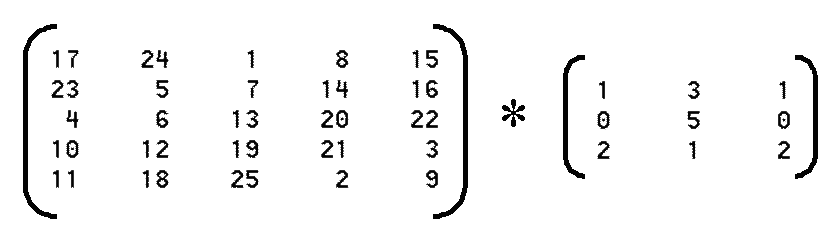
>> A=magic(5)
A =
17 24 1 8 15
23 5 7 14 16
4 6 13 20 22
10 12 19 21 3
11 18 25 2 9
>> B=[1 3 1; 0 5 0; 2 1 2]
B =
1 3 1
0 5 0
2 1 2
>> C1 = conv2(A,B,'full')
C1 =
17 75 90 35 40 53 15
23 159 165 45 105 137 16
38 198 120 165 205 197 52
56 95 160 200 245 184 35
19 117 190 255 235 106 53
20 89 160 210 75 90 6
22 47 90 65 70 13 18
>> C2 = conv2(A,B,'same')
C2 =
159 165 45 105 137
198 120 165 205 197
95 160 200 245 184
117 190 255 235 106
89 160 210 75 90
>> C3 = conv2(A,B,'valid')
C3 =
120 165 205
160 200 245
190 255 235
Since convolution is a neigborhood operation, the question arises of how to handle pixels that have no neighbors, namely those near the edge or borders of the image. There is no absolutely right way to handle this problem. Following are some suggestions based on common practices.
Make the image big enough so that there is a guard ring around the region of interest. That is, use only the valid portion of the convolution. The resulting image will be smaller by the width of the convolution operator, as suggested by the outlined region below:

Assume the image is embedded in a sea of zeros, as shown below. This
is the assumption made by the full and same methods
of conv2.

Assume that the pixels just outside the image are the same as their
nearest neighbor inside the image. This is the situation portrayed
below (obtained with add_border(rgb,'dup',width))

Consider the extended image to be a tiled version of the original, and then convolve the central image using portions of the adjacent tiles at the borders.
The following matlab code constructs a tiled image:
>> out = repmat(srgb,3,3);

A cyclic border can also be constructed from add_border(rgb,'circ',width).

One could contemplate using a fancy extrapolation scheme, perhaps based on cubic splines, to obtain pixels outside the original image. This really has no more justifcation than the other methods described above, and is no substitute for making the original image much larger than the region of interest.
function out = add_border(inp,type,width)
%
% add border of given type and width to
% an image
if (nargin<3)
width = 10;
end
if (nargin<2)
type = 'zero';
end
s = size(inp);
switch type(1)
case 'z',
s(2)=width;
z = zeros(s);
tmp = [z, inp, z];
s = size(tmp);
s(1)=width;
z = zeros(s);
out = [z;tmp;z];
case 'd',
c = inp(:,1,:);
left = repmat(c,1,width);
c = inp(:,s(2),:);
right = repmat(c,1,width);
tmp = [left, inp, right];
s = size(tmp);
r = tmp(1,:,:);
top = repmat(r,width,1);
r = tmp(s(1),:,:);
bot = repmat(r,width,1);
out = [top;tmp;bot];
case 'c',
left = inp(:,1:width,:);
right = inp(:,(s(2)-width+1):s(2),:);
tmp = [right,inp,left];
s = size(tmp);
top = tmp(1:width,:,:);
bot = tmp((s(1)-width+1):s(1),:,:);
out = [bot;tmp;top];
otherwise,
error('Invalid input arguments.');
end
Maintained by John Loomis, last updated 14 Feb 2001